Built for precision
clinical teams.
Precision Manifolds
Access high-definition single-cell baselines to benchmark clinical trajectories with 99.8% fidelity.
Evidence Registry
Full provenance tracking for research use only (RUO) applications, ensuring institutional-grade audit trails.
Simulation Lab
Deploy virtual trials to plan and manage complex reprogramming protocols with clarity and control.
OSK Partial Mode
Computationally constrained partial reprogramming — oncogene blacklist, dedifferentiation ceiling, Sirtuin scoring.
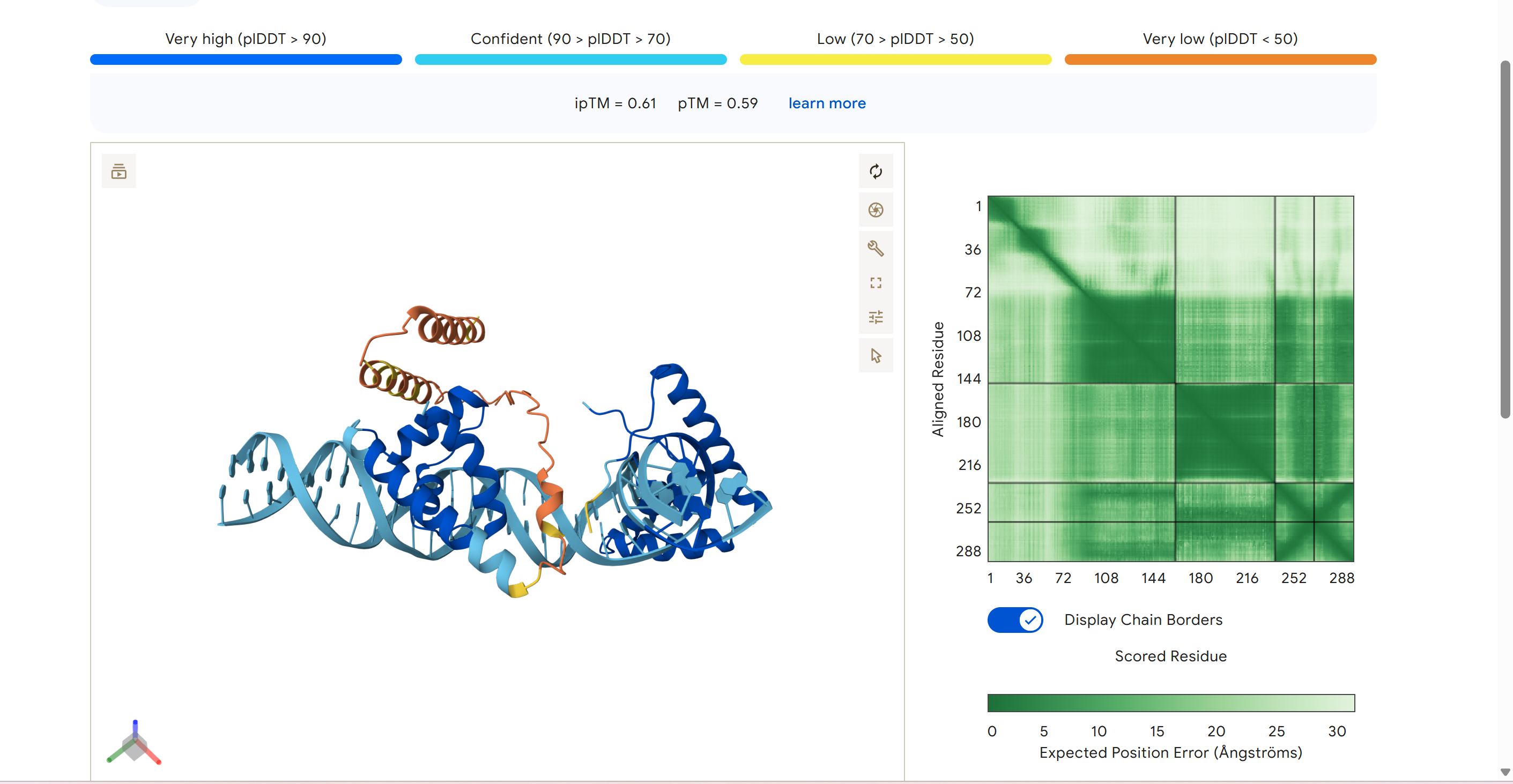
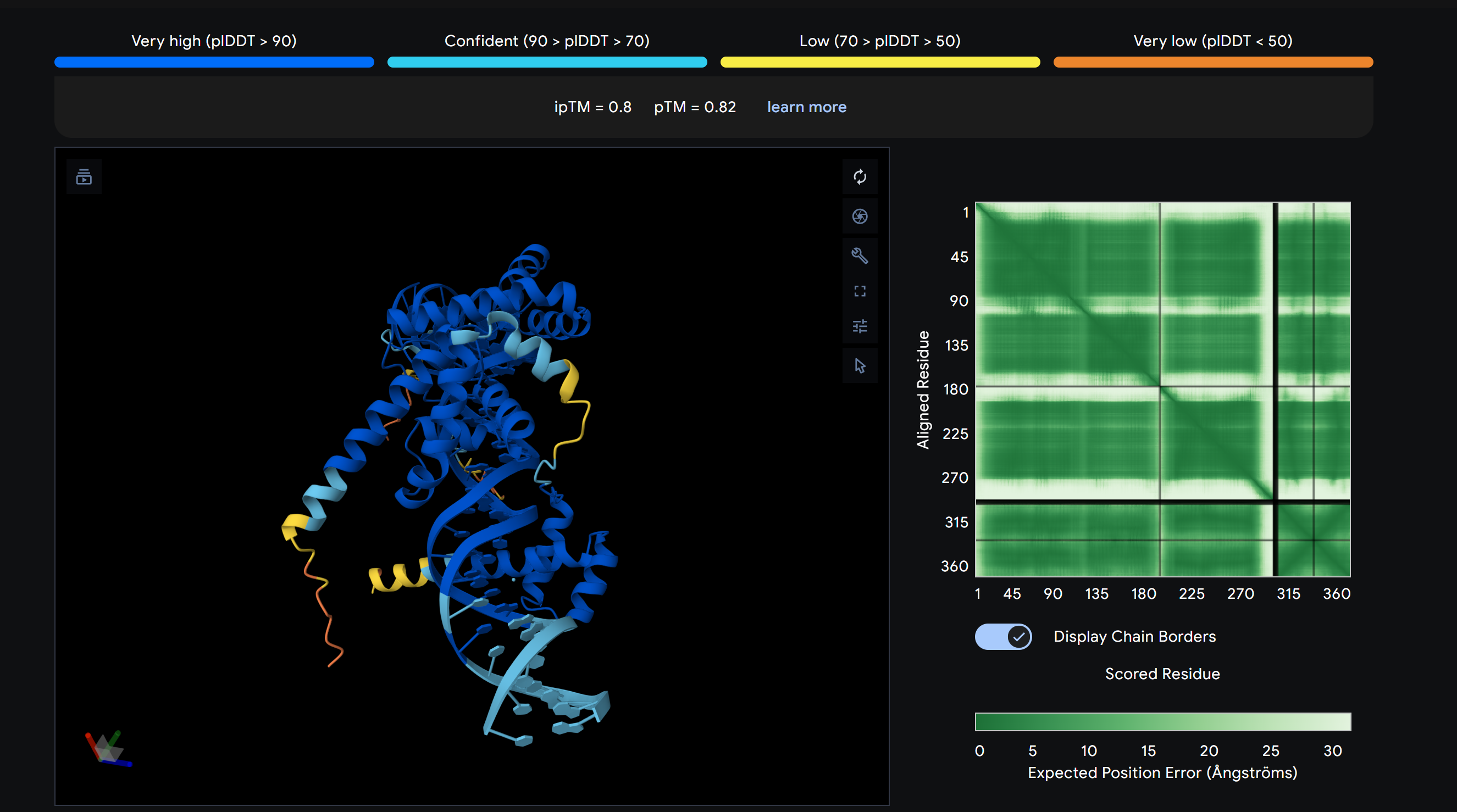
Achieving Multi-Fidelity
Structural Validation.
In this institutional benchmark, Zenith v26.4 isolated the core pluripotency 'Handshake' complex. By anchoring POU5F1 and SOX2 onto a synthetic 35bp DNA pillar, the system achieved a 0.82 ipTM score, establishing the Gold Standard for provable biological rejuvenation.
Everything your
team needs.

Propelling Life Science with
Generative
Trajectory Modeling.
The Zenith Mainframe delivers high-precision modeling for cellular longevity and reprogramming. By mapping thousands of gene dimensions, we provide the computational evidence needed to reverse biological aging.
99.8%
Model Fidelity
32 vCPUs
Compute Velocity
Autonomous Structural Authority
"Moving from manual protein-picking to a unified, autonomous structural discovery engine."
Epicardial-to-Myocardial Transformation
"Discovery of the conversion of adult epicardial cells to a myocardial fate via Elite Domain-Handshake Linkers (DHL)."
Confidence
Scaffold
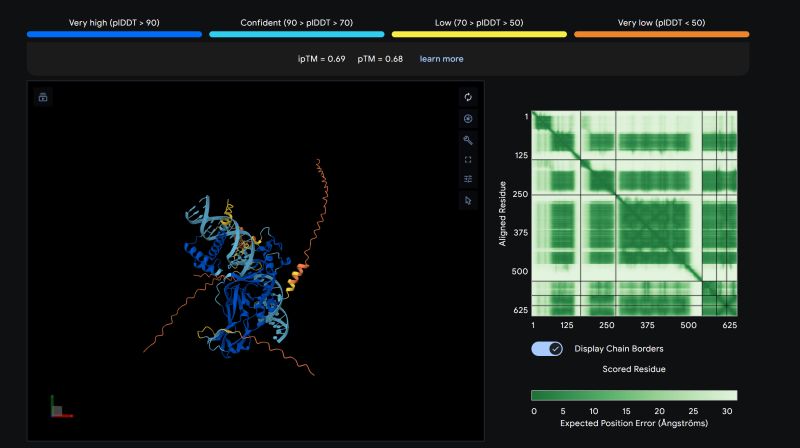
High-Fidelity Rejuvenation Manifold
AI Instruction: "Reverse biological age by 12 years. Strictly maintain ESI above 95%."
Target Reset
ESI Floor